Managing User Databases
This section is only relevant if you choose to use the built-in Instant JChem database-based security implementation or the database-based authentication implementation. This is typically useful for a multi-user database where you need to have a relatively high level of security. It is recommended for all but the simplest multi-user databases.
To be able to edit the multi-user database you must have the role ROLE_ADMIN. Without this the only operation you can perform is to change your own password. With this role you can create and delete users and change their passwords and details for them.
Step 1: Find the Security Settings editor in the schema editor.
Step 2: User management settings are located on the second tab, entitled User Management. This tab is only be present if you have specified database-based security.
Step 3: You can add and delete users, and change their attributes, including updating their password and roles using the appropriate functionality.
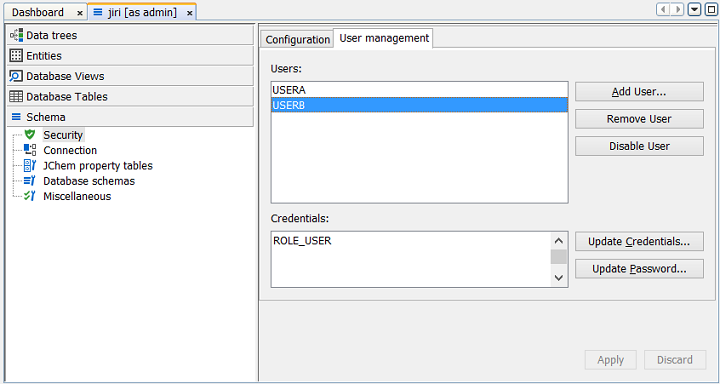
Step 4: Click Close when finished.